Simulations
Protocols for State-of-the-art Molecular Dynamics Simulations
Simulate Biological Systems with Best-in-Class Tools
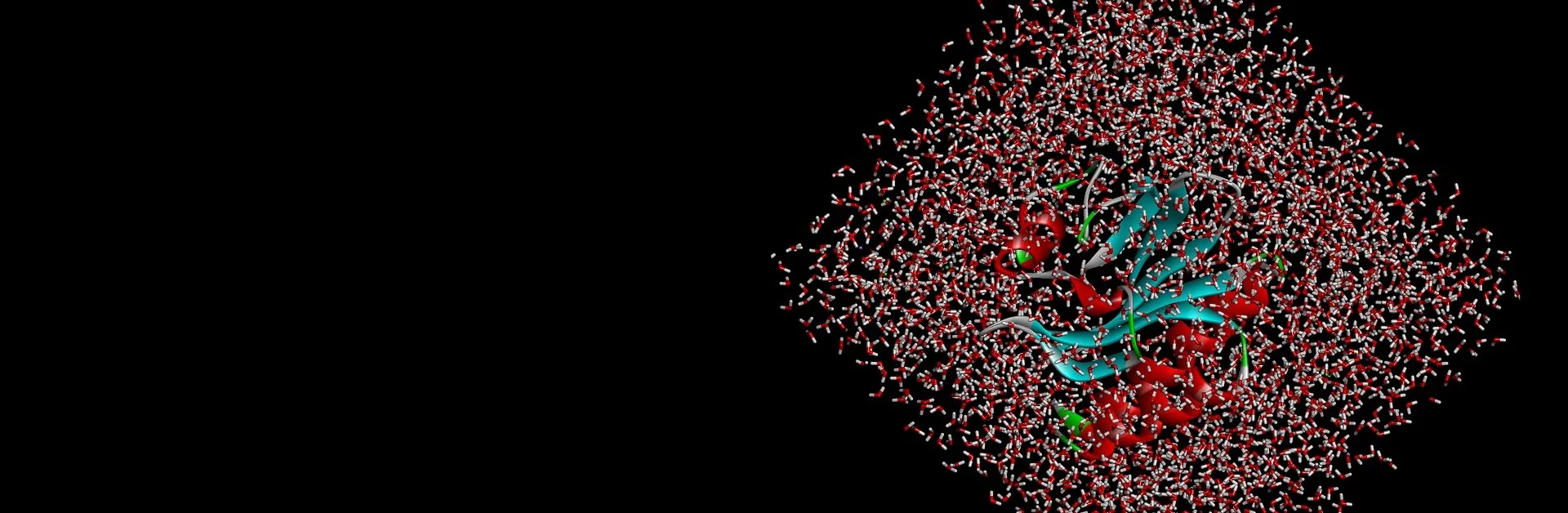
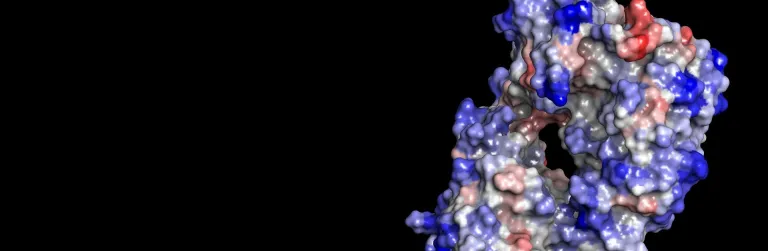
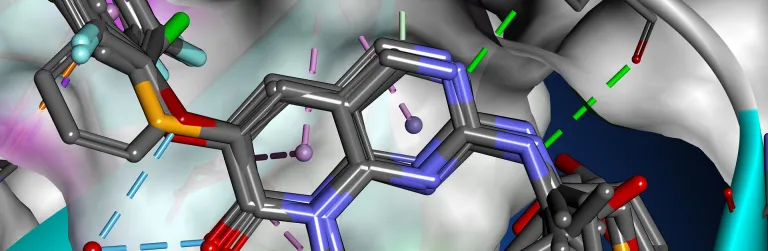
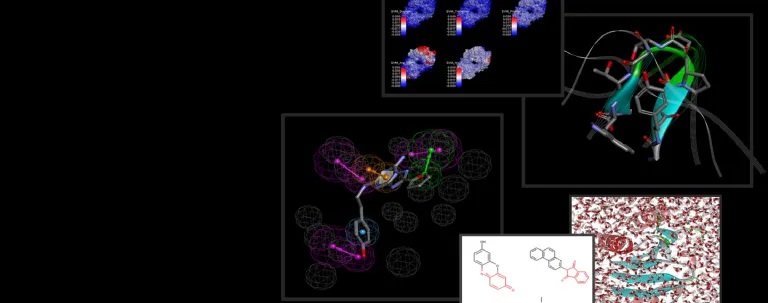
Biomolecular processes rely on a variety of dynamic interactions between proteins, ligands, solvents and ions. Often, the specifics of these interactions are difficult to capture via physical experimentation alone due to the short time scales over which they occur. Simulation can help elucidate the energetics of these processes, providing insight into their mechanism of action and properties.
BIOVIA Discovery Studio utilizes best in-class molecular simulation programs, NAMD and CHARMm. Furthermore, Gaussian accelerated Molecular Dynamics (GaMD) is also implemented in the latest release of Discovery Studio for simultaneous unconstrained enhanced sampling and free energy calculations.
- Simulate
- Model
- Explore
Simulate
- CHARMm
- NAMD
- Perform explicit solvent MD simulations
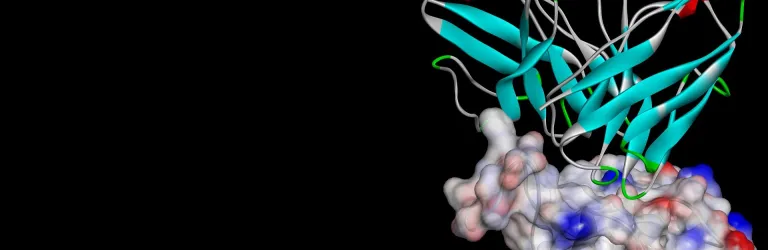
- Solvate a protein with explicit membrane and run MD simulations
- DMol3 / CHARMm
- Calculate single point energies or perform minimizations of receptor-ligand complexes using hybrid Quantum Mechanics/Molecular Mechanics (QM/MM) simulations
- Implementation of GaMD for simultaneous unconstrained enhanced sampling and free energy calculations
- Configure and run a GaMD equilibration, automatically parametrizing the boost potentials needed
- Run and restart GaMD simulations
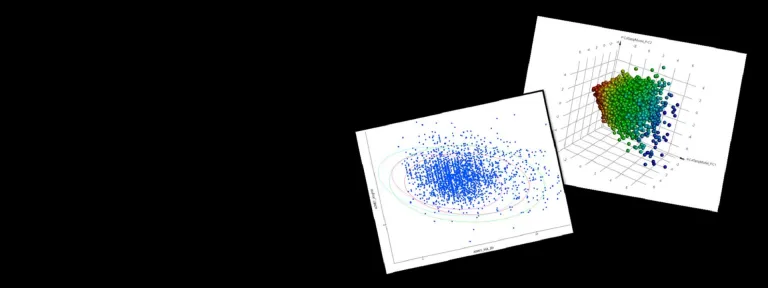
- Estimate a free energy landscape from a set of MD trajectories, allowing for statistical reweighting of GaMD simulations
Model
- Support for a broad range of force fields, including CGenFF, charmm36, CHARMm and more
- MATCH method for typing ligands with charmm36
- Full support of CHARMM patching mechanism
- Fast explicit aqueous solvation method with optional counter-ions suitable for very large molecular systems
- Solvation of transmembrane protein into pre-equilibrated lipid bilayer
- Analysis of MD trajectories
Explore
- Perform quick and accurate protein ionization and residue pKs predictions for protein preparation
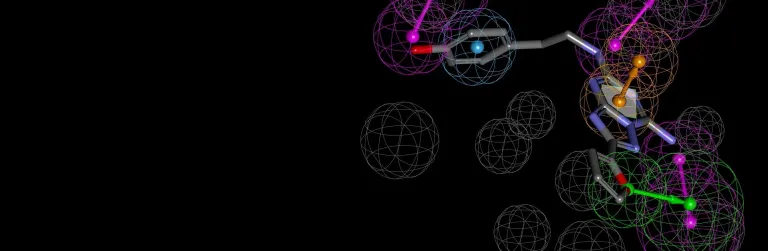
- Use CDOCKER, a CHARMm-based docking engine to perform flexible ligand-based docking and refinement
- Perform pose optimization of multiple ligands in the context of a receptor
- Calculate binding energies of docked poses
- Accurately predict relative ligand binding energy for a congeneric ligand series using the free energy perturbation (FEP) method
- Calculate the relative free energy of binding for a combinatorial library of ligands modeled by Multi-Site Lambda Dynamics (MSLD)
- Estimate ligand binding free energy and study ligand unbinding using CHARMm-based Steered Molecular Dynamics (SMD) simulations
- Examine electrostatic potential effects with CHARMm Poisson-Boltzmann (PB) equation
Start Your Journey
Accelerate your drug discovery with BIOVIA Discovery Studio.
Join the conversation in the BIOVIA Drug Discovery & Development Community!
FAQ About Molecular Dynamics Software & Programs
Also Discover
Learn What BIOVIA Can Do for You
Speak with a BIOVIA expert to learn how our solutions enable seamless collaboration and sustainable innovation at organizations of every size.
Get Started
Courses and classes are available for students, academia, professionals and companies. Find the right BIOVIA training for you.
Get Help
Find information on software & hardware certification, software downloads, user documentation, support contact and services offering